DNA Sequencing Techniques
This lesson covers:
- The development of DNA sequencing techniques
- The key processes involved in sequencing DNA
- Next-generation sequencing
The development of DNA sequencing techniques
DNA sequencing is the process of determining the exact sequence of nucleotides within a DNA molecule.
In the 1970s, Frederick Sanger and colleagues developed techniques for sequencing nucleic acids from viruses and bacteria. Sanger sequencing, which relies on the radioactive labelling of bases and a process called gel electrophoresis, was instrumental in determining DNA sequences.
The techniques he developed have undergone significant improvements, including:
- The substitution of radioactive labels with fluorescent tags for safety and efficiency.
- Enhancements in scaling up and automation to process more samples at once.
- The introduction of capillary sequencing, a key method later used in the Human Genome Project.
The Human Genome Project, an international project to map the entire human genome and share the data publicly, used these advanced sequencing and computing techniques to achieve its goals.
The processes involved in sequencing DNA
Although modern DNA sequencing methods vary, they share some fundamental principles.
A basic outline of a common sequencing approach includes:
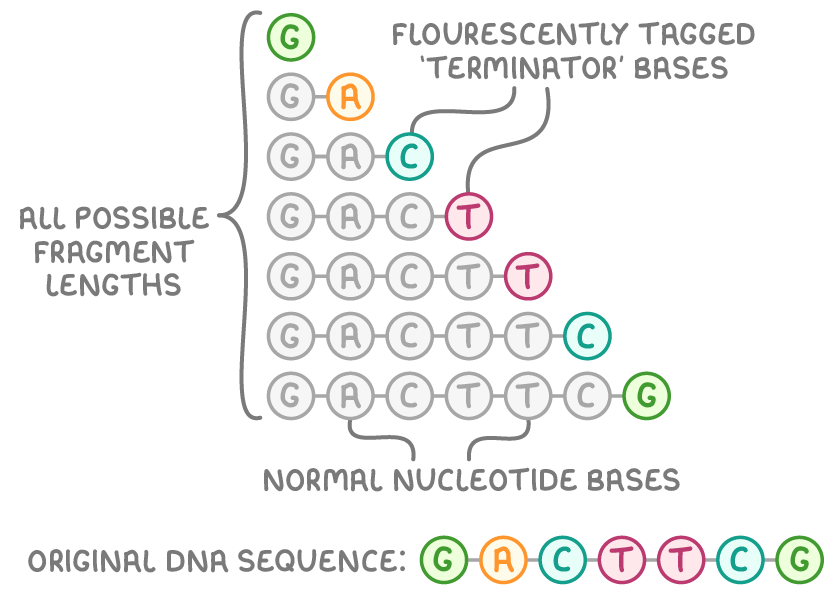
- DNA is mixed with primers, DNA polymerase, normal nucleotide bases, and 'terminator' bases.
- Double-stranded DNA is split into single strands, and copied multiple times.
- DNA polymerase adds nucleotides to the single-stranded template to start rebuilding new DNA strands.
- When a terminator base is added DNA synthesis stops, and each is tagged with a unique fluorescent colour.
- This produces DNA fragments of all possible lengths.
- These DNA fragments are separated by length, for instance using capillary sequencing or gel electrophoresis.
- A laser then detects the fluorescent colours of the terminator bases in each fragment to determine their sequence order.
- With every potential base marked, computer software analyses the fragments to reconstruct the original DNA sequence.
Computers are able to reassemble genomes by comparing and overlapping DNA fragments.
Next-generation sequencing
The early methods of Sanger sequencing were time-consuming and laborious. Next-generation sequencing refers to the automated, high-throughput technologies that have revolutionised DNA sequencing.
These new technologies have facilitated:
- Massively parallel sequencing - This allows simultaneous sequencing of millions of DNA fragments.
- Exponentially increased speed - For example, a bacterial genome can now be sequenced in less than 24 hours.
- Reduced costs - This has made it possible to sequence the genomes of more organisms.
DNA sequencing, particularly next-generation sequencing, plays a crucial role in the accurate identification of genetic disorders and cancers, marking a significant advancement in the field of genomics.